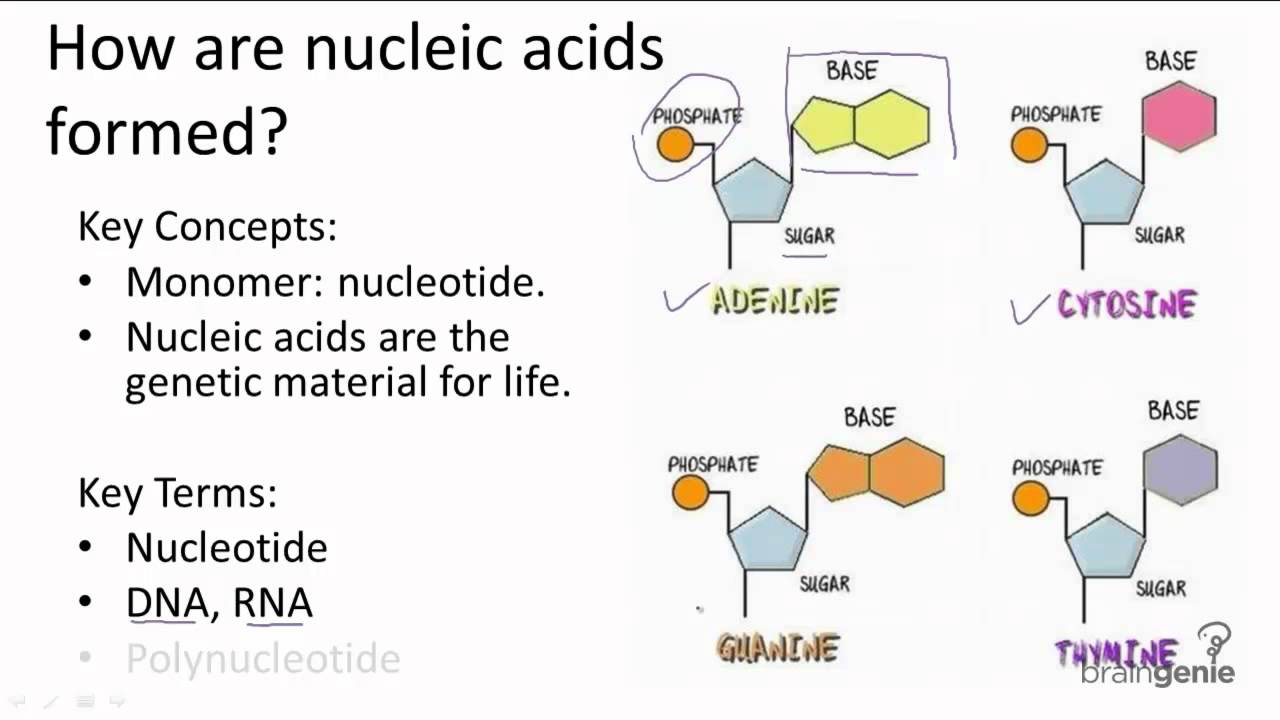
There are four types of nucleotides distinguished by which of the four nucleobases they contain: in DNA these are adenine (A), cytosine (C), guanine (G), and thymine (T). The structure of nucleic acids consists of a sequence of nucleotides. Nucleic acid double helices will only form between two strands of complementary sequences, where the bases are matched into only A-T or G-C pairs. Fundamental concepts Ĭhemical structure of DNA. In structure prediction, the structure is determined from a known sequence, while in nucleic acid design, a sequence is generated which will form a desired structure. Nucleic acid design can be considered the inverse of nucleic acid structure prediction. However, nucleic acid structures are less versatile than proteins in their functionality. Computational models for protein folding require tertiary structure information whereas nucleic acid design can operate largely on the level of secondary structure. However, nucleic acid design has the advantage of being a much computationally simpler problem, since the simplicity of Watson-Crick base pairing rules leads to simple heuristic methods which yield experimentally robust designs. 
Nucleic acid design has similar goals to protein design: in both, the sequence of monomers is rationally designed to favor the desired folded or associated structure and to disfavor alternate structures. In addition, there are many tertiary structure considerations which affect the choice of a secondary structure for a given design. 
It is necessary because there are many possible sequences of nucleic acid strands that will fold into a given secondary structure, but many of these sequences will have undesired additional interactions which must be avoided. Nucleic acid design is central to the fields of DNA nanotechnology and DNA computing. Nucleic acid design is the process of generating a set of nucleic acid base sequences that will associate into a desired conformation. 
These four strands associate into this structure because it maximizes the number of correct base pairs, with A's matched to T's and C's matched to G's. Nucleic acid design can be used to create nucleic acid complexes with complicated secondary structures such as this four-arm junction.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
